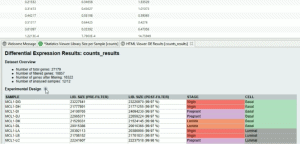
Clustering redundant transcripts in OmicsBox with CD-HIT
De novo transcriptome assemblies are required to analyze RNA-seq data from a species for which there is no reference genome. However, with the advancement of next-generation sequencing technologies, the amount of available sequencing data is growing exponentially. Because of this, assembly algorithms often generate a large number of transcripts. Removing redundancy from such data could be crucial for reducing storage space,