
Whole Genome Functional Annotation of Solanum lycopersicum
Whole genome functional annotation of Solanum lycopersicum with OmicsBox functional annotation module.

Whole genome functional annotation of Solanum lycopersicum with OmicsBox functional annotation module.
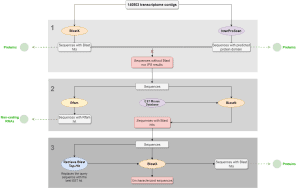
Analysis workflow Objective: To describe the process of a transcriptome characterization using Blast2GO. Input data: A mouse RNA-seq dataset composed of 140803 contigs. Pipeline: Blastx and InterproScan were performed with the complete dataset to identify proteins. Sequences with no Blastx hits nor IPS results were selected and further analyzed. RFAM and Local Blast (against an EST mouse db) were performed with the sequences with
OmicsBox allows combining two Functional Anlaysis projects which contain either have the same identifiers or different ones.

Blast2GO Supported Project. Researchers: MSc Sara D. Cardoso, Ph.D. candidate at Gulbenkian Institute for Science (IGC), Oeiras, Portugal Supervisors: Prof. Rui F. Oliveira, Gulbenkian Institute for Science (IGC), Oeiras, Portugal and Prof. Adelino V. M. Canário, CCMAR – Centre of Marine Sciences, University of Algarve, Faro, Portugal Background and Project Overview: The peacock blenny Salaria pavo (family Blenniidae) is a

BioBam Supported Project: Changes in maize transcriptome in response to Maize Iranian Mosaic Virus (MIMV) infection. Researchers: Mr. Abozar Ghorbani, Ph.D. candidate at Plant Virology Research Center, College of Agriculture, Shiraz University, Shiraz, Iran Supervisors: Prof. Keramatollah Izadpanah, Plant Virology Research Center, College of Agriculture, Shiraz University, Shiraz, Iran and A/Prof. Ralf Dietzgen, Queensland Alliance for Agriculture and Food Innovation, the
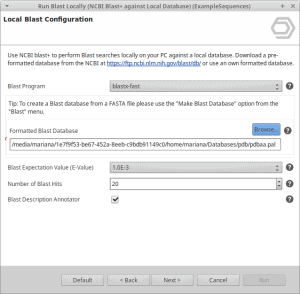
OmicsBox allows running Blast locally. The blast algorithm will run on the user’s computer against a database that is installed locally. In order to do so, we have to either download a pre-formatted NCBI database or format our own database (see this tutorial until step 3). Pre-formatted databases can be downloaded directly from the NCBI ftp or via a Perl script provided by
(Release Date: 01/10/2015) You can now perform Blast and InterProScan from within the CLI, which means you can run the whole functional annotation analysis with just one command. Find here some useful command line examples.

