We are happy to announce OmicsBox Web, a new browser-based companion to BioBam’s OmicsBox platform. OmicsBox Web is available at https://omicsbox.biobam.com and uses the same OmicsBox account credentials, so analysis jobs launched via OmicsBox, input data and results as well as account settings are now reachable from any device with a web browser.
OmicsBox Web is designed to complement the desktop application, not to replace it. The desktop remains the primary environment for configuring and running bioinformatics analyses. What OmicsBox Web adds is a lighter path to the day-to-day operations that do not require the full interface: checking on running cloud jobs, moving files in and out of cloud storage, opening result files, and managing the subscription. For more detail on the interface and each page, see the OmicsBox Web section of the user manual.
What OmicsBox Web covers
After sign-in, OmicsBox Web is organised in five pages: Dashboard, Files, Jobs, Results, and Settings. Each one mirrors a piece of the desktop application and is designed to feel familiar to existing users.
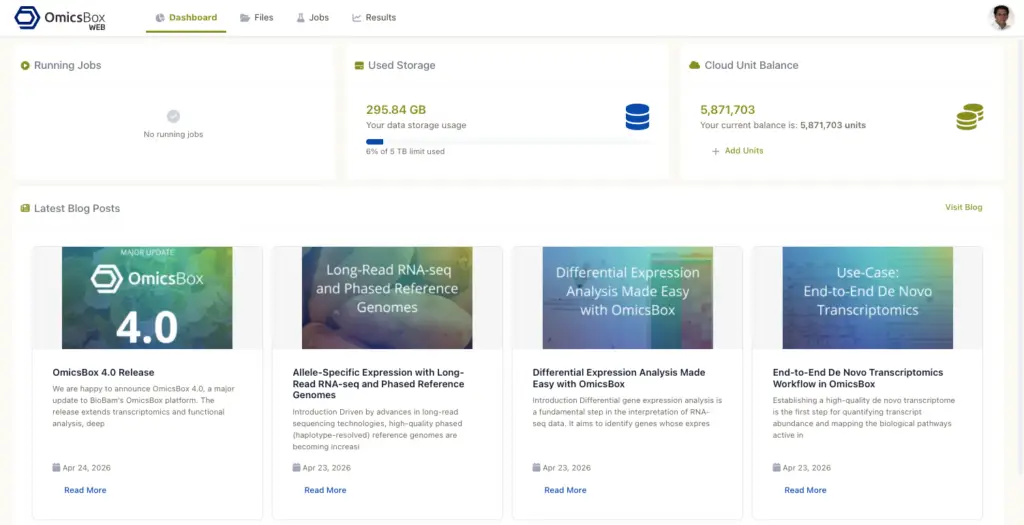
Dashboard
The Dashboard is the landing page. It shows three summary cards: Running Jobs with name, ID, and elapsed time; Used Storage with a progress bar against the cloud storage quota; and Cloud Unit Balance with a link to recharge. Clicking on a job opens the full Jobs page. The Dashboard also lists the most recent entries from the BioBam blog, so new features, tutorials, and bioinformatics resources are easy to find.
Files
The Files page is a browser-based file manager for the user’s cloud storage. It offers the same functionality as the Cloud Files view in the desktop application: browse folders and files, upload by drag-and-drop or with the upload button, download to the local computer, create or delete folders, and open .box files in the Results viewer.
The Files page is also the place to bring in input data for analyses. Uploaded files become immediately available to Cloud Sync tools both from the desktop application and from OmicsBox Web, which makes it practical to prepare datasets on one device and launch the work from another. For background on cloud storage, see Cloud Files in the manual.
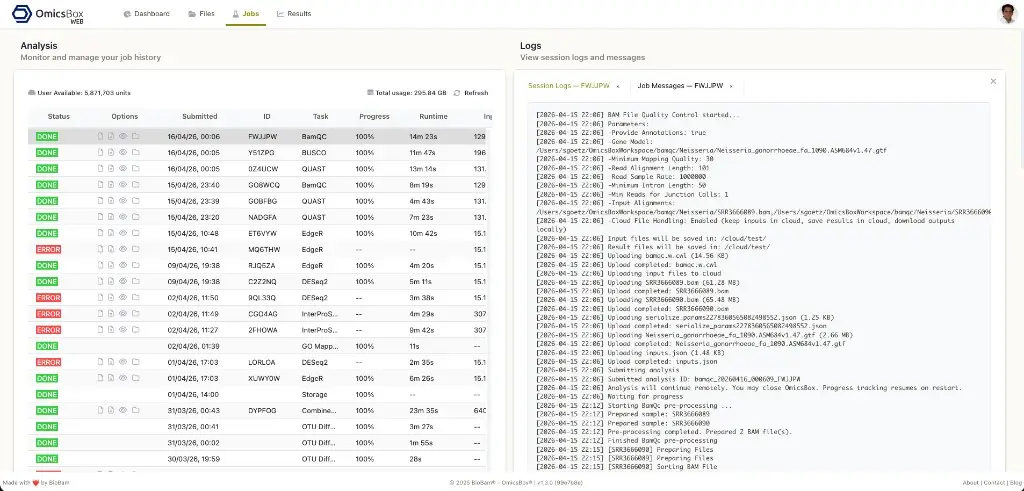
Jobs
The Jobs page lists the cloud job history in a detailed table. It uses the same Cloud Usage component as the desktop application, so the columns, actions, and filters are identical: status, progress, runtime, log viewer, message viewer, and job details. Running jobs can be cancelled from this page, and the table can be filtered and sorted by any column, including date ranges.
In addition to the desktop view, the Jobs page in OmicsBox Web uses a side panel layout. Selecting the log or details icon opens the corresponding panel next to the job table, so job metadata, parameters, and algorithm output are visible without leaving the page.
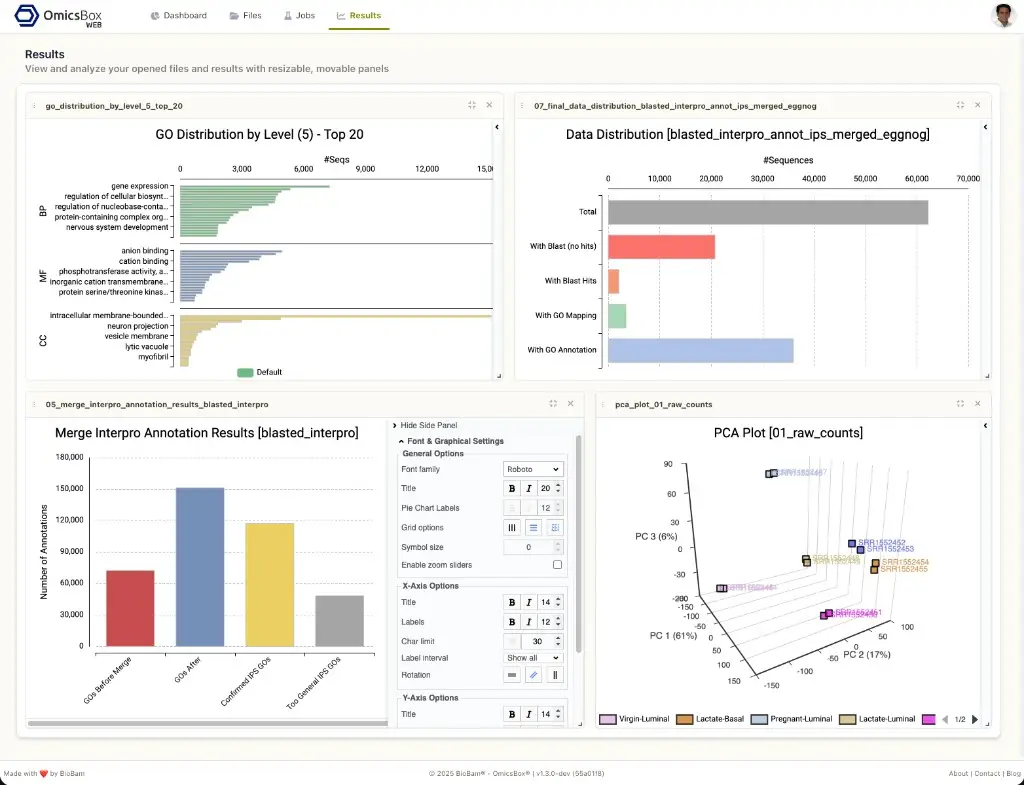
Results
The Results page provides an interactive viewer for analysis output files. When a .box file is opened from the Files page, it is displayed in Results using resizable and movable panels. Several result files can be open at the same time, each in its own panel, and the layout is automatically saved between sessions. The supported visualisations include charts, pathway diagrams, and other outputs produced by OmicsBox tools, such as GO distributions, annotation results, and PCA plots.
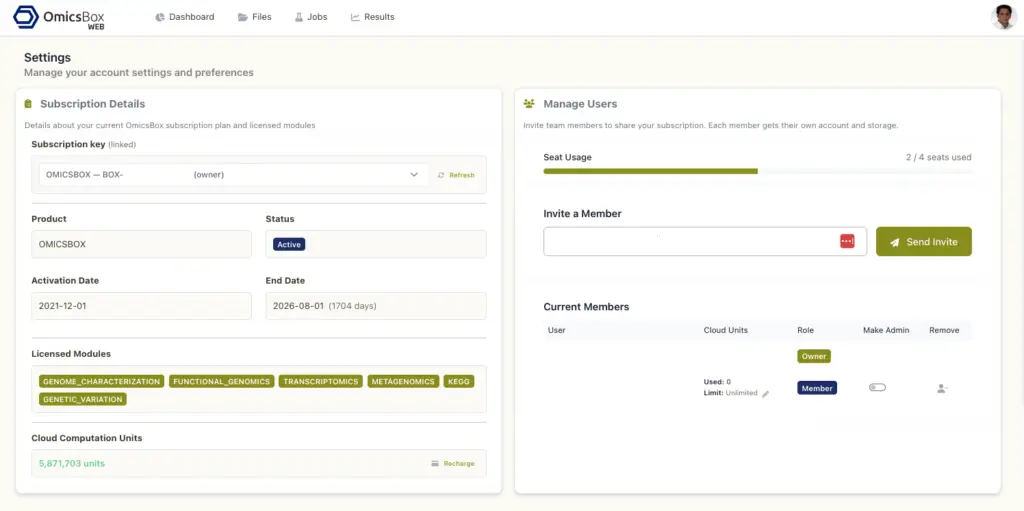
Settings
The Settings page collects account management and configuration options. Subscription details show the product name, status, activation and end dates, licensed modules, subscription key, and Cloud Computation Units balance. Administrators can add or remove authorised users from the User management section, which is useful for shared team subscriptions. The Security and session section includes SMS two-factor authentication and a preference for how long the user stays signed in on each device.
Getting started
OmicsBox Web is live at omicsbox.biobam.com. Existing users can sign in with their OmicsBox email and password, or with Google Sign-In if the account is linked. No installation or extra licence is required; the service is part of the standard OmicsBox subscription. Step-by-step descriptions of each page are in the OmicsBox user manual.
About OmicsBox
OmicsBox is BioBam’s bioinformatics software for the analysis of omics data. It provides workflows for genome, transcriptome, and metagenome datasets, organised in five modules: Genome analysis, Transcriptomics, Genetic Variation, Functional analysis, and Metagenomics.
The Functional analysis module is based on the Blast2GO annotation methodology, with more than 10,000 citations in the scientific literature, and is widely used in non-model organism research.
Computationally demanding steps, such as genome assemblies and other CPU- or memory-intensive pipelines, are executed in the cloud. OmicsBox itself runs on a standard PC or laptop under Windows, Linux, or macOS. Licensing is subscription-based, with flexible options.
Feedback, questions, and feature requests are welcome at support@biobam.com or through the BioBam account. Further information, documentation, and contact options are available at biobam.com/omicsbox and in the user manual. Trade names: BioBam, OmicsBox, and Blast2GO as used by BioBam.
About the Author
 Dr. Stefan Götz
Dr. Stefan GötzStefan is the founder and CEO of BioBam. Since founding the company in 2011, he has focused on helping researchers simplify genomics and omics data analysis through practical and accessible software solutions. Under his leadership, BioBam has supported scientists and institutions worldwide in advancing research in human health, agriculture, and environmental science. Stefan remains closely involved in product strategy, innovation, and customer-driven development. He holds a Ph.D. in bioinformatics and combines scientific expertise with entrepreneurial leadership to make advanced bioinformatics more accessible to the global research community.

