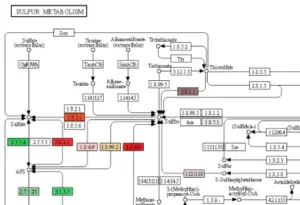
Transcriptome analysis reveals important metabolic pathways of Allium sativum
OmicsBox was used to perform the complete transcriptome analysis of Allium sativum. Enrichment analysis revealed important metabolic pathways.
Product Tutorial, Quickstarts, New Features, etc.

OmicsBox was used to perform the complete transcriptome analysis of Allium sativum. Enrichment analysis revealed important metabolic pathways.
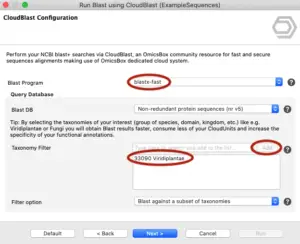
Using cloud resources efficiently is crucial for researchers dealing with large datasets. This article will show you how to use CloudBlast effectively, helping you save on computational costs and avoid common pitfalls. OmicsBox allows you to perform NCBI Blast searches using a cloud system for high-performance computing. This system runs jobs in parallel and autoscales depending on demand. To control

This project shows how OmicsBox was used to functional annotate Moringa oleifera and to better understand the genome of the plant.

Understanding the molecular differences that underlie the different COVID19 progression severity with Omic

OmicsBox Supported Project. Researchers: Hayden Waller, Cornell University, USA Background and Project Overview: One of the primary outcomes of speciation – the formation of new species – is the generation of pre-mating barriers that prevent or greatly reduce gene flow. Behavioral changes are perhaps the most potent in the early stages of speciation. In particular, the divergent evolution of sexual

De novo transcriptome assemblies are required to analyze RNA-seq data from a species for which there is no reference genome. However, with the advancement of next-generation sequencing technologies, the amount of available sequencing data is growing exponentially. Because of this, assembly algorithms often generate a large number of transcripts. Removing redundancy from such data could be crucial for reducing storage space,
Third-generation DNA sequencing technologies allows scientist to generate longer sequence reads, which can be used in whole-genome sequencing projects to yield better repeat resolution and more contiguous genome assemblies. However, although long-read sequencing technologies can produce genomes with long contiguity, the relatively high error rate of long reads has made it challenging to generate highly accurate final sequences. OmicsBox now

De novo transcriptome analysis in Datura metel and identification of genes related to withanolides biosynthesis pathway Researchers: Madhavi Hewadikaram Dr Sanjaya Deepal Bathige Prof. Veranja Karunaratne Project Overview Withanolides are secondary plant compounds that belong to a family of C28 ergostane-type steroidal δ lactones that mainly belong to the family Solanaceae of the plant kingdom. Withanolides have held interest and

BioBam Supported Project – with OmicsBox. Researchers: Dr. Abhishek Kumar, Computational Genome Biology Lab at Institute of Bioinformatics, Bangalore, India Project: The pandemic of coronavirus disease 2019 (COVID-19) caused by the novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pushed the entire world on a halt with the largest lockdown of our times. A total numbers of infected cases are
