How to create a taxonomic mapping file.
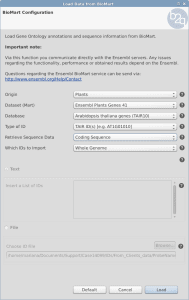
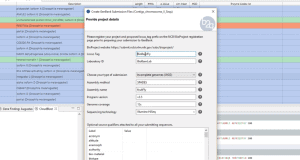
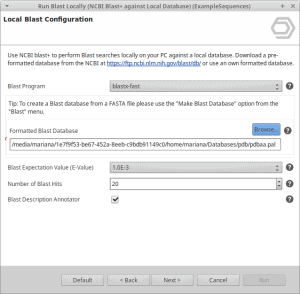
Create a taxonomic mapping file to Make Blast Database within OmicsBox OmicsBox allows creating a custom database to run Blast locally. The blast algorithm will run on the user’s computer against a database that is installed locally.In order to do so, we have to either download a pre-formatted NCBI database (see tutorial) or format our own database (see this tutorial