OmicsBox enables users to rename all selected sequences by modifying the existing sequence name through conversion, replacement, or adding text.
Use Cases
1. Rename sequence suffixes using regular expressions
It is possible that someone noticed too late that the sequence IDs in the Blast results and in the FASTA file used for InterProScan are slightly different.
The prefix of the FASTA file is similar, but there is an extra extension in the last part of the sequence name.
Example:
| Blast results | InterProScan |
| seq1 | seq1_(ORF) |
| seq2 | seq2_(ORF) |
| seq3 | seq3_(ORF) |
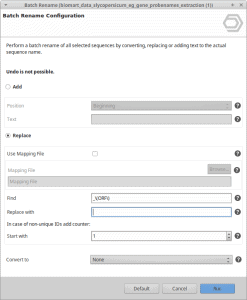
OmicsBox offers the so-called Batch Rename feature, which uses the regular expression to search for a term in the sequence name and change it to the desired one.
This feature is available in the Functional Analysis module and can be found in the side panel in the Tools section.
Using the data from the example, there are 2 different solutions.
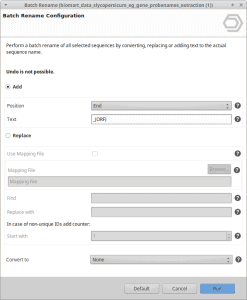
Solution 1: Add the extra extension to the blast results project.
- The suffix to be added is _(ORF).
Figure 1: Add the term to the end of the sequence name.
Now that the identifiers match, combining both projects and adding the Fasta and the InterProScan results to the project with the blast results is possible.
To combine both projects visit the following Tips & Tricks.
2. Rename Using a Mapping File
The batch rename feature also allows you to use a mapping file to perform the sequence name replacement. This mapping file should contain the original sequence names you want to rename and their replacements, separated in 2 columns by a tab.
In this use case, we have a tomato dataset with non-informative sequence names, and we want to replace these names with the original gene IDs from Ensembl. The mapping file used for this purpose looks like this (Figure 3).
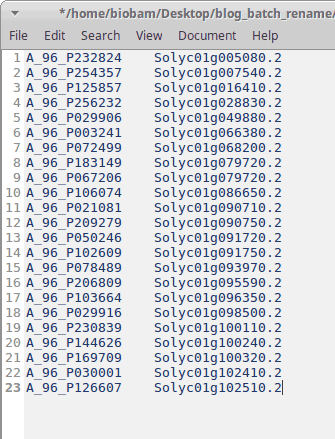
Figure 3: Mapping file.
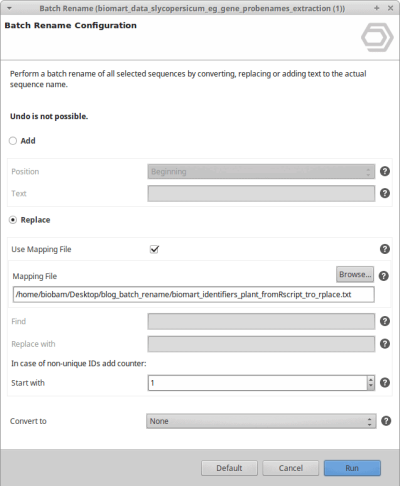
We can do the following:
Use the mapping file to rename the original sequence names.
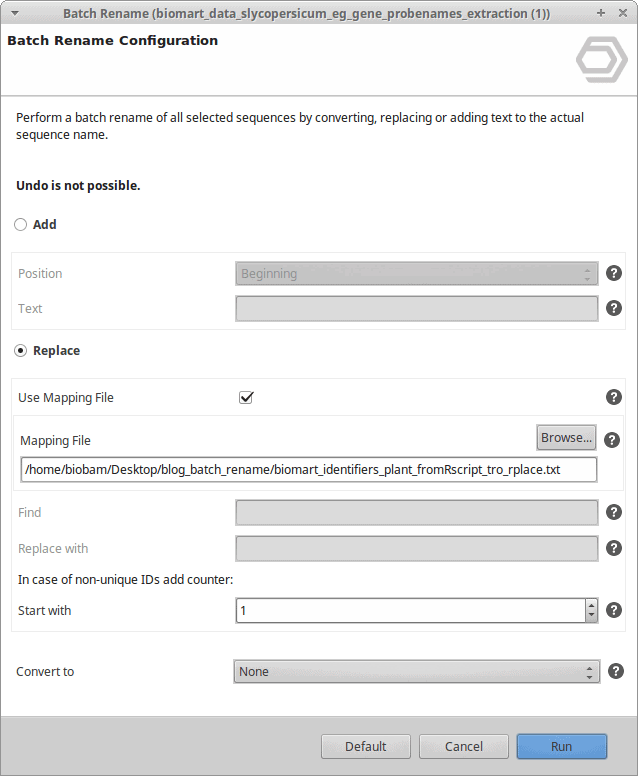
After running this process, the sequence names have been replaced and now contain the gene identifiers instead.
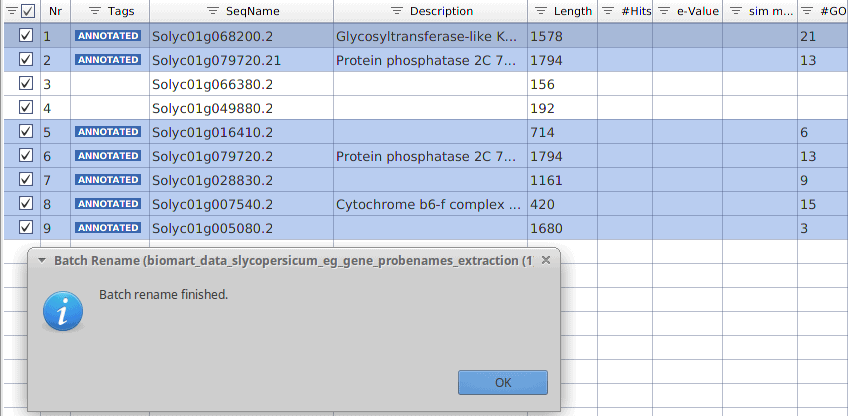
Figure 4: Sequence names replaced with the Ensembl gene IDs.
This blog has been updated with OmicsBox 3.2. information