Blast2GO Supported Project.
Researchers:
- Prof. Luis Delaye, CINVESTAV Irapuato, México
- Prof. Enrique Calderon, Instituto de Biomedicina de Sevilla, España
- Prof. Andrés Moya, I2SysBio, Universidad de Valencia, España
Background and Project Overview:
Fungi from the genus Pneumocystis parasites lungs of mammals. These fungi show extensive stenoxenism, meaning that each Pneumocystis specie parasites a single mammal species. In humans, Pneumocystis causes pneumonia in immunocompromised hosts. Pneumocystis genomes are endowed with several copies of a gene coding for a Major Surface Glycoprotein (MSG). Pneumocystis uses MSG proteins for host attachment and immune system evasion.
Three Pneumocystis genomes have been sequenced. These are the genomes from Pneumocystis jirovecii, Pneumocystis carinii and Pneumocystis murina. The species infecting humans, rats and mice, respectively. The objective of this work is to study the pattern of natural selection on Pneumocystis genomes.
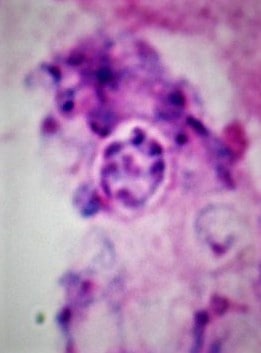
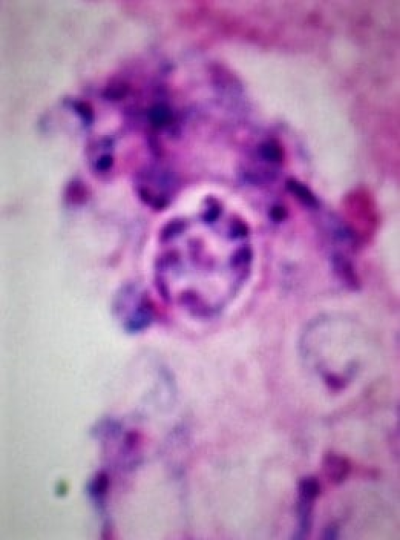
Cyst from Pneumocystis showing eight endospores. Giemsa stain.
Contribution of Blast2GO:
Blast2GO was used to identify functionally related sets of genes showing accelerated rates of evolution. To do so, we used Gene Set Enrichment Analysis (GSEA) within Blast2GO. In addition, we used Directed Acyclic Graphs (DAG) to explore the results from GSEA. These powerful tools allowed us to discover that genes involved in secretion of MSG proteins show accelerated rates of molecular evolution.
Research Centers:
CINVESTAV Irapuato is a public research center devoted to the study of microorganism and plant biotechnology. Is located in the state of Guanajuato, México.
Instituto de Biomedicina de Sevilla is a multidisciplinary center devoted to biomedical research in Spain.
I2SysBio is a research center from the University of Valencia devoted to study the structure, function, dynamics, evolution, and manipulation of complex biological systems.