Coding Potential Assessment Tool
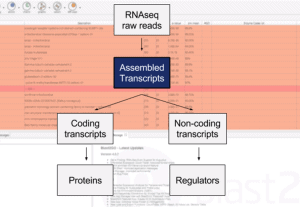
This video shows how to use the ‘Coding-Potential Assessment Tool’ which allows distinguishing the coding transcripts from the non-coding transcripts. This can be achieved using prebuilt models or building a species-specific model from the NCBI database. The results of the coding potential can be cross-checked with the Blast results and may allow discovering some novel mRNA.



