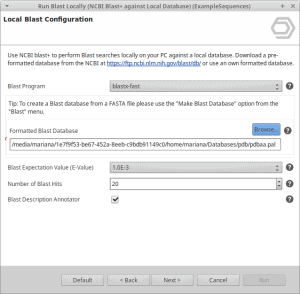
Error when running LocalBlast “NCBI Binaries C++ Exception”.
The error looks like: “Critical: (110.6) CNcbiRegistry: Syntax error in system-wide configuration file: NCBI C++ Exception: “……….\src\corelib\ncbireg.cpp”, line 660: Error: ncbi::IRWRegistry ::x_Read() – Badly placed ‘\’ in the registry value: ‘ROOT=J:\nASNLOAD=J:\BioEdi t\tables\nDATA=J:\BioEdit\tables\’ (m_Pos = 4)Error: NCBI C++ Exception: “……….\src\corelib\ncbireg.cpp”, line 660: Error: ncbi::IRWRegistry ::x_Read() – Badly placed ‘\’ in the registry value: ‘ROOT=J:\nASNLOAD=J:\BioEdi t\tables\nDATA=J:\BioEdit\tables\’ (m_Pos = 4)” This error is