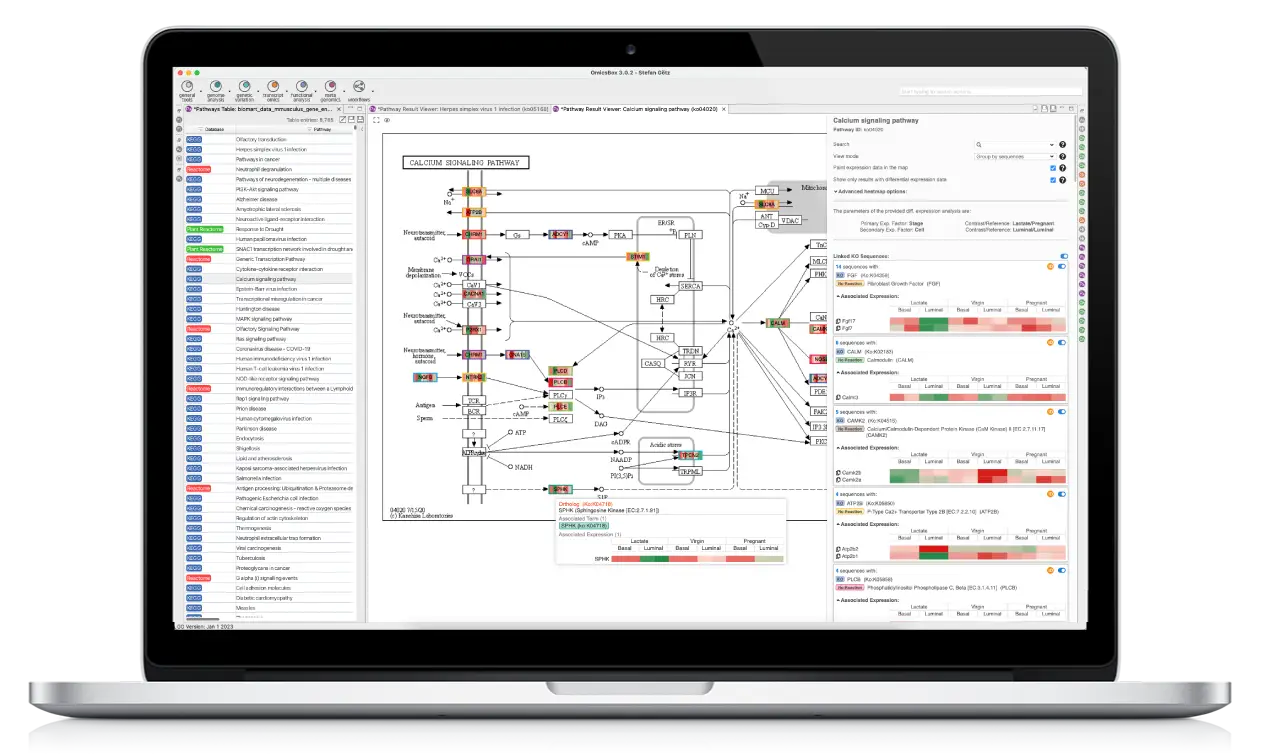
Your All-In-One Bioinformatics Software Solution

Over 15k researchers choose OmicsBox for their projects!
OmicsBox is the leading bioinformatics solution that offers end-to-end NGS data analysis of genomes, transcriptomes, and metagenomes.
We have designed OmicsBox to be user-friendly, efficient, and with a powerful set of tools to extract biological insights from omics data.
OmicsBox is used by top private and public research institutions worldwide. It allows researchers to easily process large and complex data sets, and streamline their analysis process.
Five modules to easily process large and complex data sets
From raw reads to functional insights fast and easy
Easily convert raw DNA-Seq reads into a structurally annotated and curated draft genome.
Identify and analyze genetic variations within a population or a species.
Allows different types of microbiome analysis like assembly and taxonomic classification.
Over 500 Institutions around the world trust OmicsBox










Take your research to the next level with OmicsBox
Get to Know Us

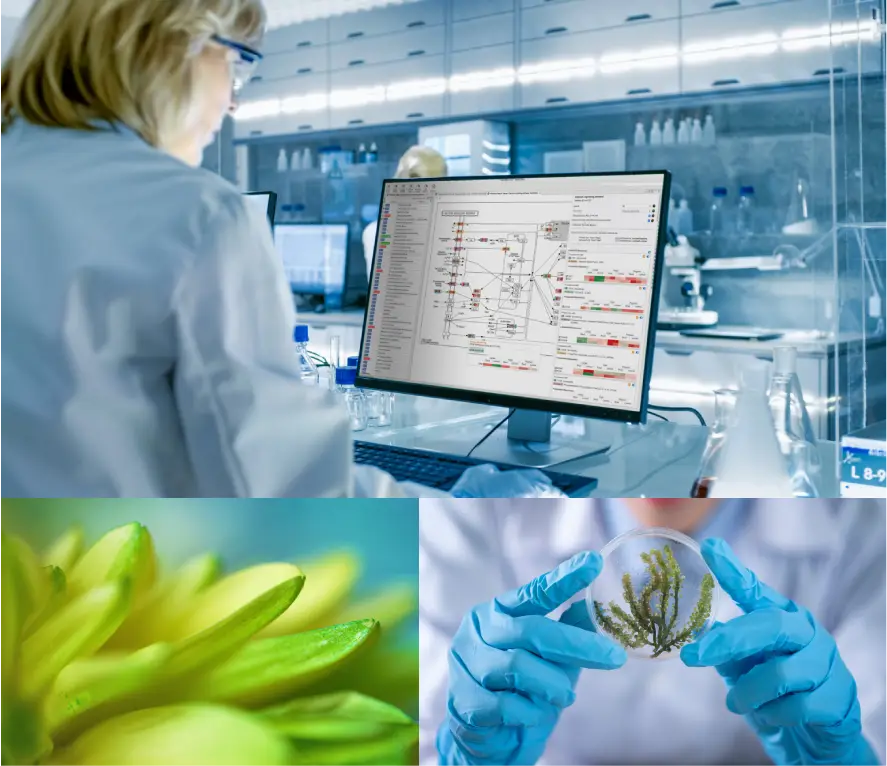
BioBam is a leading bioinformatics solution provider which accelerates research in disciplines such as agricultural genomics, microbiology, and environmental NGS studies, amongst others.
At BioBam we are committed to the development of user-friendly software solutions for biological research. Our mission is to transform complex data analysis procedures into attractive and interactive tasks. Our ultimate goal is to close the gap between experimental work, bioinformatics analysis, and applied research.
Latest Blog Articles


OmicsBox 3.2 Release
Paula Cubilles
April 12, 2024

Using BWA for DNA and RNA Alignment in OmicsBox
Enrique Presa
March 7, 2024

Differences Between GTF and GFF Files in Genomic Data Analysis
Stefan Götz
February 27, 2024

Tips to manage my OmicsBox subscription users
Stefan Götz
February 26, 2024